Their SMART method accelerates enzyme evolution by reducing the selection period for superior variants from several weeks to a few days, and decreases overall enzyme engineering campaign costs by eliminating the need for specialized equipment.
Enzymes are proteins that catalyze chemical reactions in living organisms. They are widely applied in industries such as food production, detergents, pharmaceuticals, and chemicals. However, for commercial use, natural enzymes often need improved stability, substrate specificity, or catalytic efficiency.
Directed evolution is a Nobel Prize-winning strategy for improving proteins. It introduces artificial mutations into their genes and then selects superior variants. This approach mimics natural evolution over several weeks instead of millions of years.
A significant challenge of this approach is that artificially induced mutations can generate up to 100 trillion candidate variants, which renders the screening process extremely time-consuming and expensive.
To address this challenge, researchers at Nagoya University and their colleagues have developed SMART (Single-Molecule Assay on Ribonucleic acid by Translated product), an in vitro selection platform.
Their study demonstrated that SMART identifies improved enzyme variants much more rapidly and cost-effectively than conventional methods. The findings were published in the journal ACS Synthetic Biology.
The SMART system was developed by a research group led by Associate Professor Jasmina Damnjanović and Professor Hideo Nakano of the Graduate School of Bioagricultural Sciences at Nagoya University, in collaboration with researchers from the Institute of Science Tokyo and Saitama University. This approach successfully combines mRNA display, next-generation sequencing, and bioinformatics.
Key features of the SMART system
Typically, proteins and genes are physically separate, making it difficult and time-consuming to identify which gene encodes a discovered enzyme.
In the SMART system, puromycin acts as a chemical bridge, linking the enzyme protein to its corresponding blueprint, messenger RNA (mRNA). This mRNA display technique enables precise tracking of the relationship between individual proteins and their encoding genes at the single-molecule level.
Nakano emphasized, “In principle, there is no method for enzyme screening that is more efficient than this system. Screening enzymes at the single-molecule level has rarely been attempted before.”
SMART also incorporates an auxiliary unit for detecting enzyme activity. This study used engineered ascorbate peroxidase 2 (APEX2) as the auxiliary enzyme for oxidase screening. When the target oxidase is active and releases hydrogen peroxide (H₂O₂), APEX2 attaches a biotin marker to nearby molecules, enabling their isolation and capture.
Enzyme screening experiments using SMART
The researchers chose a yeast oxidase, SpDAAO, as a model enzyme because it has great potential for drug synthesis and diagnostics. The selection prioritized D-amino acids as enzyme substrates due to their growing relevance in medical applications.
The SMART method consists of several steps—creating a DNA library of enzyme variants, synthesizing enzymes in vitro, forming an mRNA display library, labeling catalytically active enzymes, isolating them with magnetic beads, and using sequencing data to guide subsequent rounds.
To assess the method, the team tested it on a simulated library with different ratios of active and inactive variants. After a single selection round, active variants were highly enriched, confirming SMART’s effectiveness.
In practical experiments, the team generated a mutant library by substituting the essential 232nd amino acid with each of the 20 other amino acids. Next-generation sequencing analysis showed that the wild-type (original form) Y232 was clearly selected (p < 0.001), reinforcing the method’s selectivity.
Initially, genetic analysis indicated selection of several variants, in addition to the original form. However, further statistical analysis identified these as experimental noise with minimal practical significance, supporting the method’s specificity.
Conclusion and future perspectives
The experiments showed that SMART selection is highly effective. At the same time, the team recognized the need for rigorous statistical analysis and careful experimentation, rather than relying solely on initial results.
The researchers expect SMART to be applicable beyond oxidases. They aim to facilitate the integration of novel enzymes into industry, establishing the system as a foundation for future enzyme development and practical biocatalytic solutions.
Publication
Kalhari Munaweera, Nana Odake, Hannah Patricia Halim, Kakeru Ikeda, Bo Zhu, Maurizio Camagna, Tomokazu Ito, Tetsuya Kitaguchi, Naoto Nemoto, Hideo Nakano, and Jasmina Damnjanović (2026). Harnessing the Power of SMART Single-Molecule Display for Enzyme Evolution: A Focus on Oxidase, ACS Synthetic Biology. DOI: 10.1021/acssynbio.5c00968
Funding
This work was supported by Japan Society for the Promotion of Science (JSPS) Grant-in-Aid for Early-Career scientists [grant number JP18K14387 and JP22K14828] and Grant-in-Aid for Transformative Research Areas (A) (Publicly Offered Research) [grant number JP25H02263], the Collaborative Research Program by Network Joint Research Center for Materials and Devices (Ministry of Education, Culture, Sports, Science and Technology -Japan: MEXT), and Retention, Development, and Promotion Program Program Aiming at Maximizing the Career Potential of Female Researchers, Nagoya University, (MEXT’s Initiative for Realizing Diversity in the Research Environment, Leadership training type for women) awarded to Jasmina Damnjanović, and in part by Pre-Research Unit System of the Institute of Integrated Research, Institute of Science Tokyo and JSPS Grant-in-Aid for Transformative Research Areas (A) (Publicly Offered Research) [grant number JP24H01123] awarded to Bo Zhu.
Expert contact
Jasmina Damnjanović
Graduate School of Bioagricultural Sciences, Nagoya University
Email: jasmina@agr.nagoya-u.ac.jp
Media contact
Naomi Inoue
International Communications Office, Nagoya University
Email: icomm_research@t.mail.nagoya-u.ac.jp
Top image
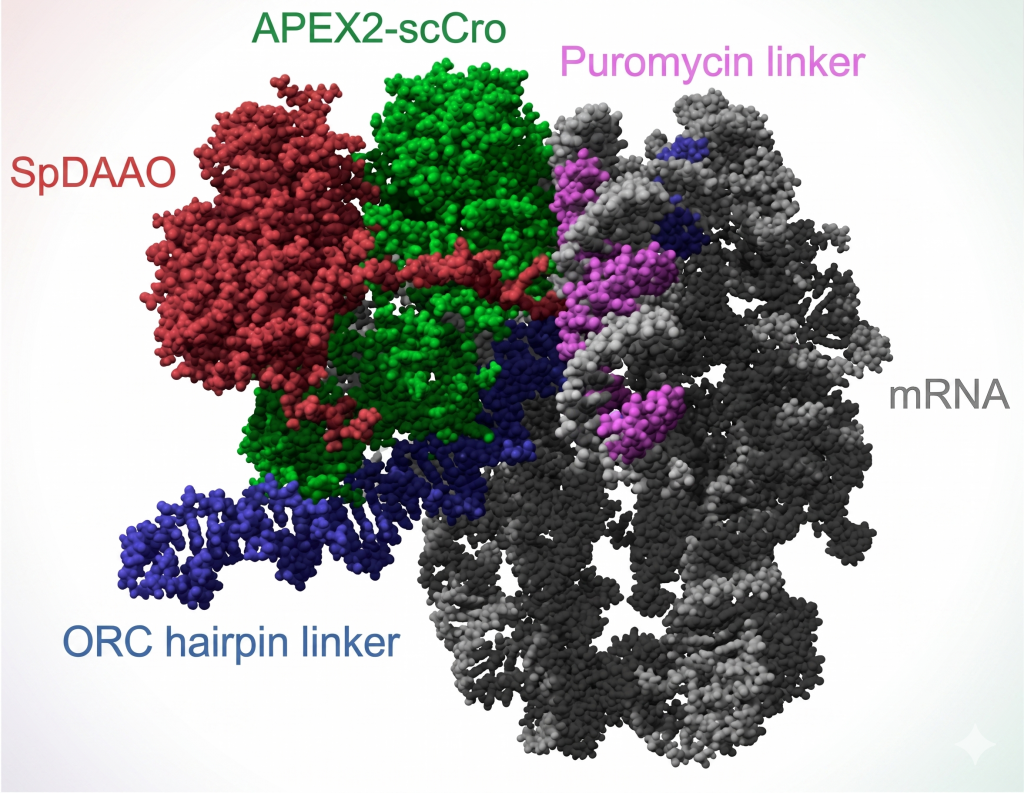
The SMART single-molecule display model, predicted by Alphafold3, shows SpDAAO (red) linked to a puromycin linker (magenta) through puromycin incorporation into the growing polypeptide. The mRNA (gray) is hybridized and chemically joined to the linker, connecting it to its protein, SpDAAO. An auxiliary unit is added using ORC hairpin DNA (blue) with APEX2-scCro fusion protein (green).
Credit: Hideo Nakano and Jasmina Damnjanović